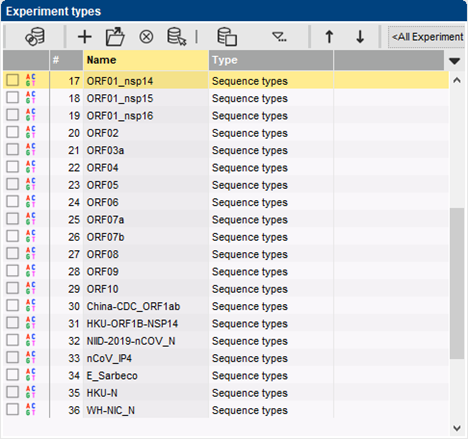
The BIONUMERICS SARS-CoV-2 plugin tool facilitates the processing and analysis of SARS-CoV-2 genomic sequences, whether downloaded from a public data repository or generated locally.
Each genomic sequence imported into BIONUMERICS is separated by the plugin tool into subsequences matching the different CDSes. Next, each of these sequences is analyzed for SNPs relative to the NCBI reference sequence for SARS-CoV-2, i.e. NC_045512.
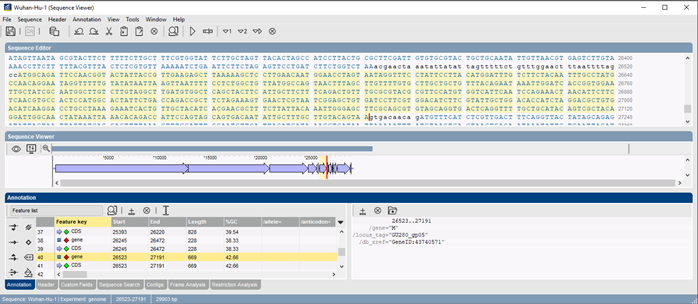
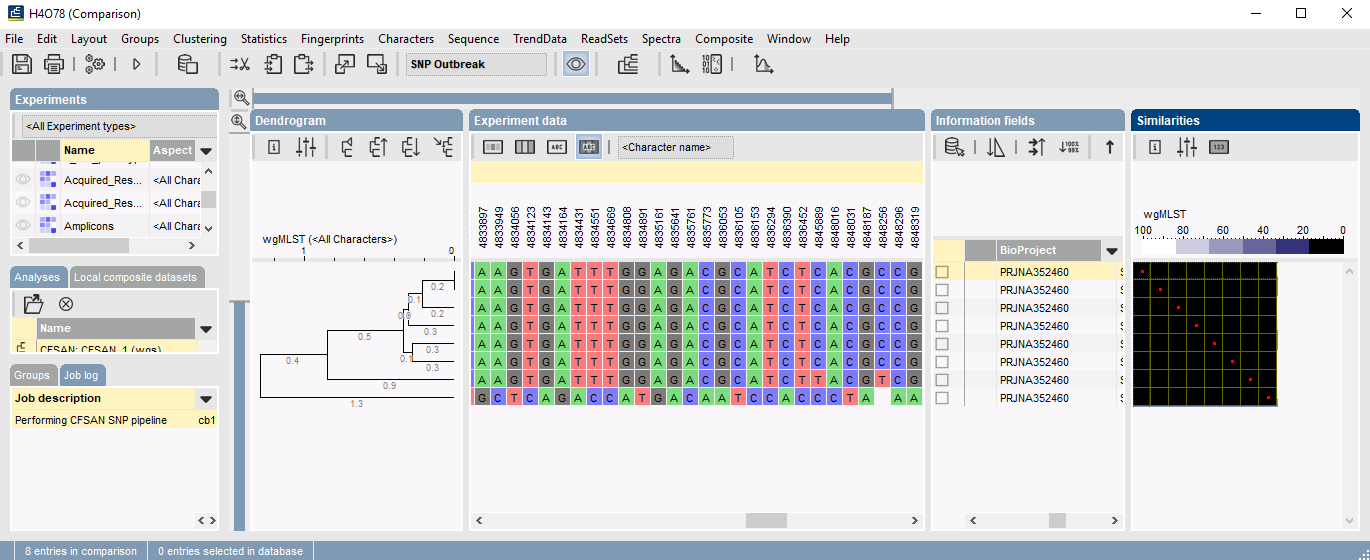
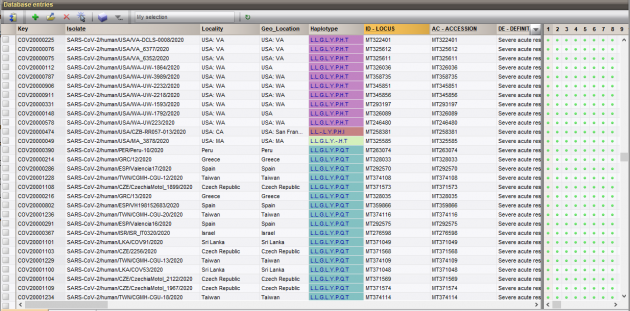
SNPs are stored in the database and can be used for easy comparison and strain typing using BIONUMERICS’ clustering tools to generate dendrograms and minimum spanning trees. Sequences are also translated, enabling SNP interpretation based on amino acid changes.
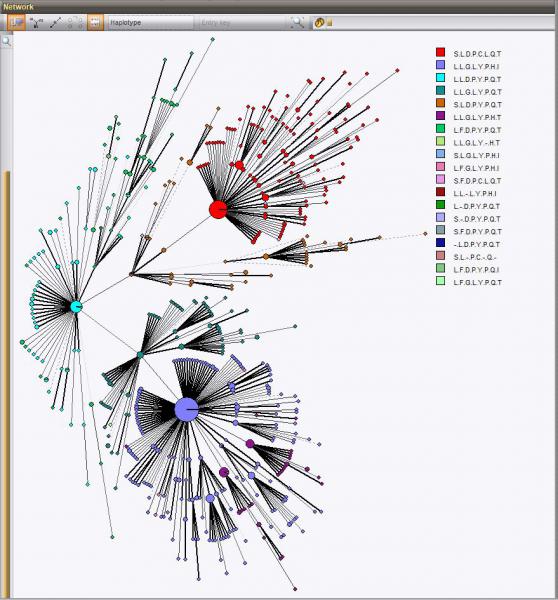
The haplotype is determined by categorization of a set of translated common missense SNPs into amino acids. This haplotype information is also stored in the database and can be displayed on the trees and networks for easy group detection.
Based on the WHO standard primer sequences, PCR products are extracted from the genome sequences and used for further in-depth analysis.